| STRuster results: 88710 |
| SCOP 1.71 classification |
| Class: | Alpha and beta proteins (a/b) |
| Fold: | TIM beta/alpha-barrel |
| Superfamily: | Cobalamin (vitamin B12)-dependent enzymes |
| Family: | Methylmalonyl-CoA mutase, N-terminal (CoA-binding) domain |
| Protein: | Methylmalonyl-CoA mutase alpha subunit, domain 1 |
| Species: | Propionibacterium freudenreichii, subsp. shermanii |
| Species SCOP sunid: | 88710 |
| Set description |
| Number of entries: | 15 |
| SCOP Entries: | d1e1ca1 d2reqa1 d7reqc1 d1reqa1 d5reqa1 d4reqa1 d7reqa1 d6reqc1 d1reqc1 d5reqc1 d4reqc1 d1e1cc1 d2reqc1 d3reqa1 d6reqa1 |
| Alignment File: | 88710.alig |
| Aligned Sequences: | 88710_alig.fasta |
| Hierarchical clustering |
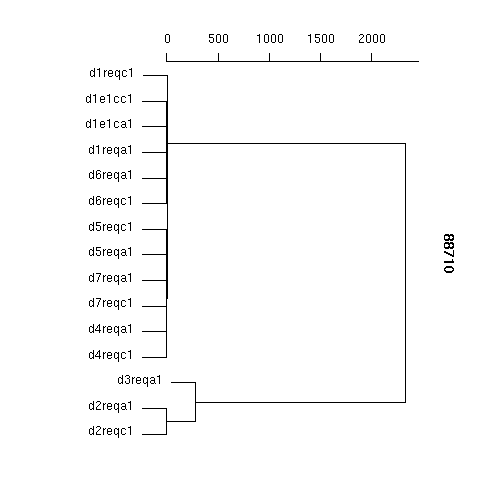
| Dissimilary matrix: | 88710_dissim_matrix.txt |
| Atom Distance Standard Deviation Plots |
| S matrix: Distance Standard Deviation |
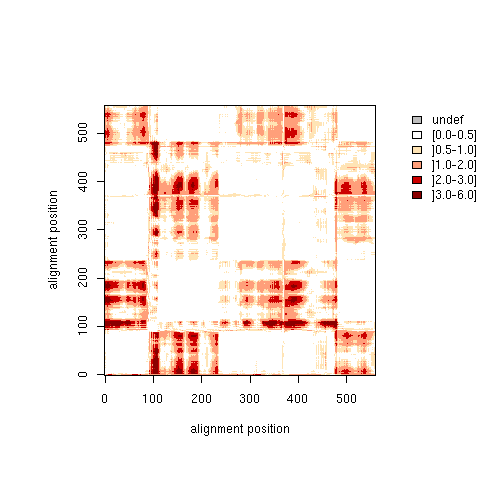
| Higher resolution plot |
| S matrix |
| T matirx: Total Standard Deviation |
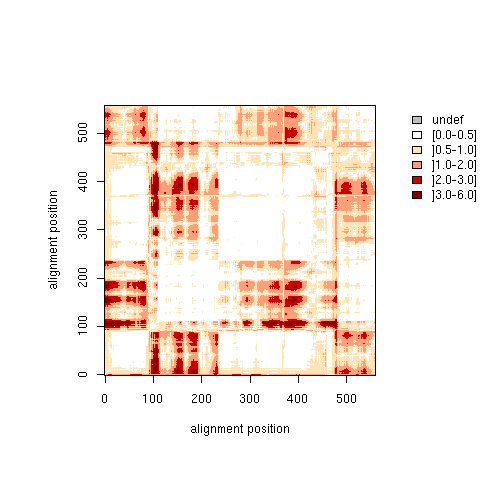
| Higher resolution plot |
| T matrix |
| R matrix: Relative Distance Standard Deviation |
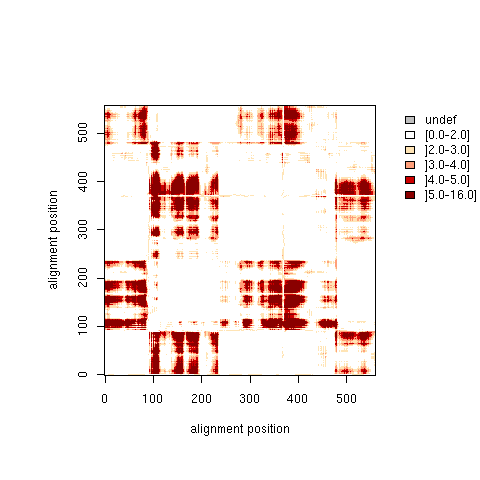
| Higher resolution plot |
| R matrix |
| Hinges and Invariant Regions (T) |
| Hinges |
| Alignment Nr | [(0, 1), (89, 96), (107, 109), (136, 138), (234, 235), (367, 369), (472, 481)] |
| Uniprot Nr | [(1, 2), (90, 97), (108, 110), (137, 139), (235, 236), (368, 370), (473, 482)] |
| Invariant Regions |
| Region | 1 |
| Alignment Nr | [(97, 106), (110, 135), (139, 233), (482, 558)] |
| Uniprot Nr | [(98, 107), (111, 136), (140, 234), (483, 559)] |
| Superposition | 88710_superp_1.pdb.gz |
| Rasmol Script | 88710_superp_1.rsm |
| View in Jmol | |
| Region | 2 |
| Alignment Nr | [(2, 88), (236, 366), (370, 471)] |
| Uniprot Nr | [(3, 89), (237, 367), (371, 472)] |
| Superposition | 88710_superp_2.pdb.gz |
| Rasmol Script | 88710_superp_2.rsm |
| View in Jmol | |
| Region | 3 |
| Alignment Nr | [(110, 135), (236, 366)] |
| Uniprot Nr | [(111, 136), (237, 367)] |
| Superposition | 88710_superp_3.pdb.gz |
| Rasmol Script | 88710_superp_3.rsm |
| View in Jmol | |